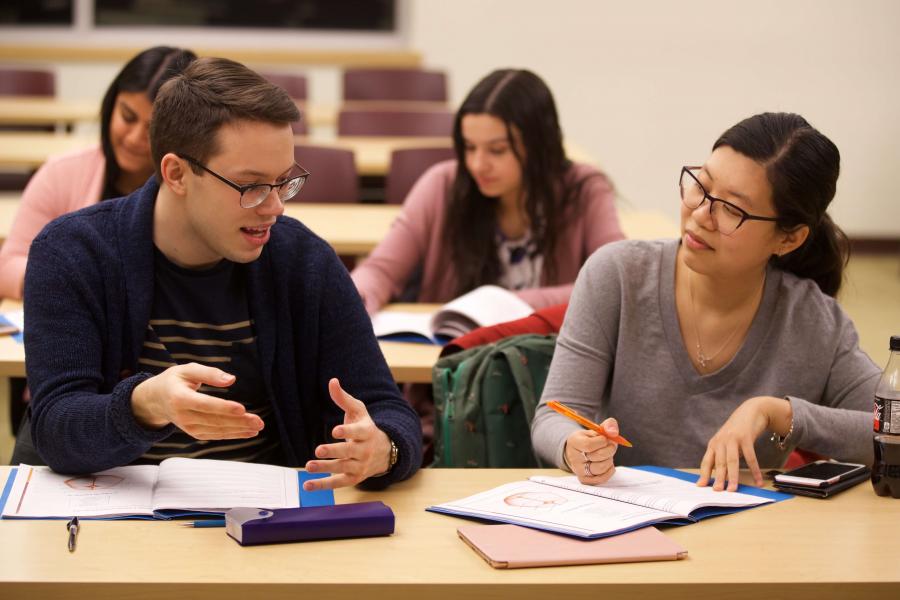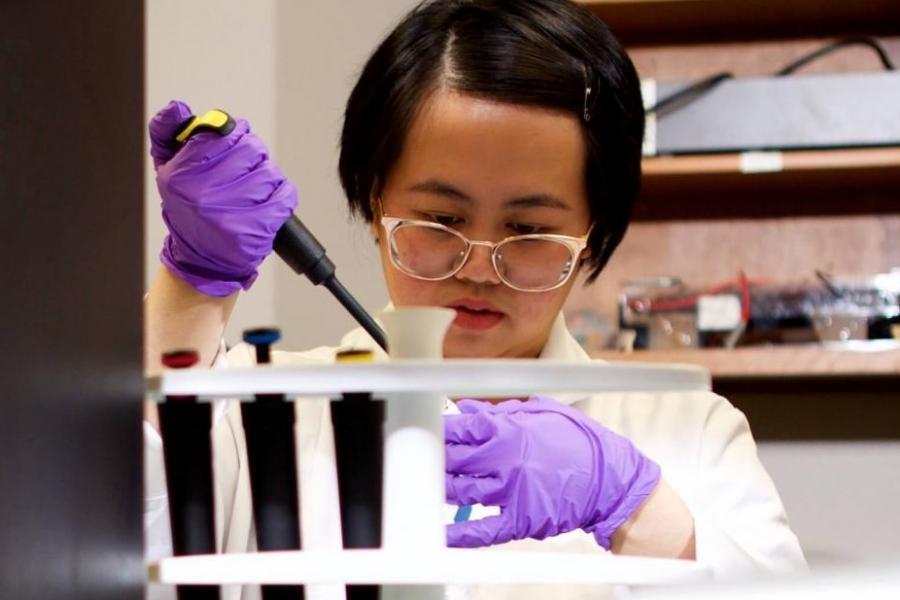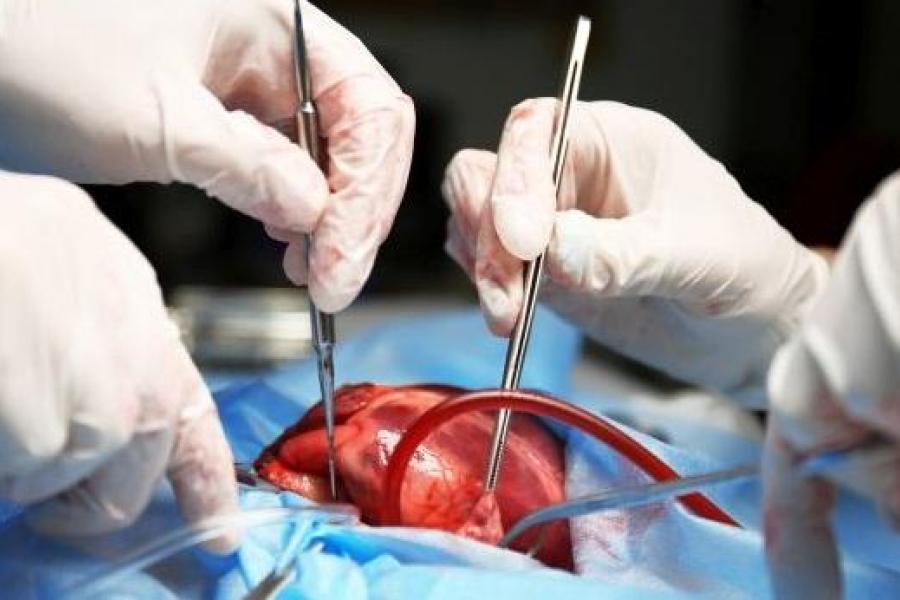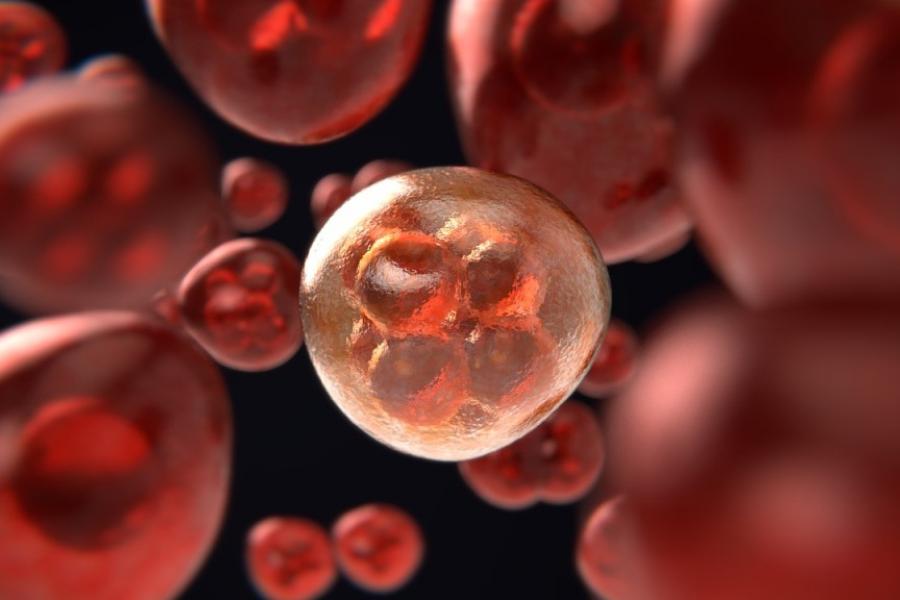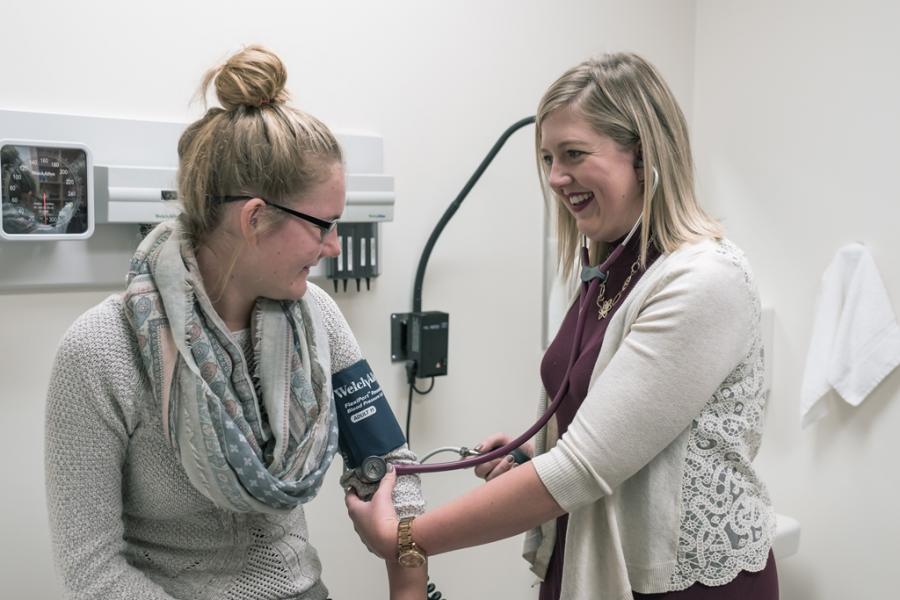
What we offer
Our department offers PhD and MSc graduate degrees in physiology and pathophysiology and a one-year stand-alone post-baccalaureate diploma in medical physiology and pathophysiology.
Our story
Watch a brief video to learn more about our department and what we offer.
Department research
Researchers in the Department of Physiology & Pathophysiology are carrying out leading-edge research across a broad range of disciplines.
Our researchers
Current faculty research
Faculty and staff
Find an advisor for your graduate studies and training
You must have an advisor willing to accept you as a student to pursue graduate studies in physiology and pathophysiology. If accepted, the advisor will commit to a minimum stipend. Currently, this is $19,000 per year for two years for a MSc degree and $20,000 per year for four years for a PhD degree, but please contact us to confirm approved level and duration of support. Trainees are all encouraged, however, to apply for personnel awards that, in addition to recognition, may provide alternative or additional support during the period of training.
If you are interested in pursuing a graduate degree under the mentorship of a faculty researcher listed on this site under links to “Our researchers”, “Faculty” or “Other faculty”, you are welcome to contact that faculty member directly to inquire about potential graduate positions.
Below, however, are links to faculty members associated with our Department that are interested in receiving trainee applications at this time. Please note, while every effort is made to update this information regularly, circumstances may have changed since the last update.
When you decide to contact a faculty member regarding a graduate position, please send them the following:
- Curriculum vitae or resume
- All academic transcripts from previous post-secondary institutions as well as the University of Manitoba (as applicable)
- Descriptions of any previous research or teaching-related experience
Please also expect an invitation for interview, if your inquiry is pursued by a prospective advisor.
Advisors accepting graduate students
| Faculty advisor | Research area |
|---|---|
|
Spinal cord physiology; movement; neural circuit formation (MSc and/or PhD applications accepted) |
|
|
Smooth muscle cell biology, cell signaling and pharmacology; pulmonary circulation hemodynamics (MSc and/or PhD applications accepted) |
|
| Robin Da Silva | Adenosine receptors and macrophage signalling, mitochondrial function and one-carbon metabolism (MSc and/or PhD applications accepted) |
|
Clinical research focused on the measurement of intraspinal pressure and spinal cord perfusion following acute traumatic spinal cord injury in humans. These measures will also be studied in the context of biomarkers for spinal cord injury. |
|
|
Cardiac stem cell therapy and tissue engineering, Immunomodulatory biomaterials synthesis, characterization and application, allograft vasculopathy, iPSCs and disease modeling, Inborn metabolic disorders (MSc and/or PhD applications accepted) |
|
|
Cardiac fibrosis. Myofibroblast inactivation by endogenous repressor of Smads. Generation of living valve leaflets using matricryptic signals (MSc and/or PhD applications accepted) |
|
|
Studies on (1) the role of the circadian clock in the regulation of the pancreatic beta cell and insulin secretion; and (2) the mechanisms that contribute to beta cell dysfunction in offspring exposed to diabetes during gestation (MSc applications accepted) |
|
|
Pathobiology and pathogenesis of asthma, including the effects of inhaled pollutants, and pre-clinical testing of new therapies (MSc and/or PhD applications accepted) |
|
|
Translational neuroscience; neural regeneration; neural stem cell biology and therapeutics; multiple sclerosis and spinal cord injury; treatment development; neuroinflammation and immune modulation; myelin repair; organoid; transgenic and genetic models (MSc and/or PhD applications accepted; For PhD recruitment previous background in neuroscience demonstrated in publication is required.) |
|
|
Interested clinically in minimal invasive pediatric general surgery. Research is focused on congenital anomalies including congenital diaphragmatic hernia (CDH) and abnormal lung development, including delineating the role of microRNAs and circular RNAs (MSc and/or PhD applications accepted) |
|
|
Cell death; autophagy; mitochondrial metabolism; hypoxia. (MSc and/or PhD applications accepted) |
|
|
Cancer; genomic instability; nuclear architecture; imaging (MSc applications accepted) |
|
|
Basic science (animals, in vitro) and translational (humans) studies; microvascular physiology; critical care; sepsis; skeletal muscle dysfunction (MSc and/or PhD applications accepted) |
|
| Suresh Mishra | Interplay between adipose and immune functions in normal physiology, aging, and diseases, with a current emphasis on exploring the role of sex hormones, genes, and micro-RNAs in defining sex differences in these functions to improve understanding of sex-biased metabolic and immune diseases. (MSc and/or PhD applications accepted) |
|
Circadian factors and pathways by which circadian misalignment leads to cardiometabolic dysfunction; how controlling feeding/fasting cycles affect cardiac function in cardiometabolic syndrome and contribute or prevent the development of cardiometabolic heart failure (MSc and/or PhD applications accepted) |
|
|
Cardiovascular lipidomics (MSc and/or PhD applications accepted) |
|
|
Hemodynamics and biomarker identification in cardiovascular diseases (MSc and/or PhD applications accepted) |
|
|
Synapse development; synaptic plasticity; neurodevelopment; autism; schizophrenia (MSc applications accepted) |
|
|
Type 1 Diabetes; autoimmunity; pancreatic islet biology; cellular senescence; DNA damage response; ER stress (MSc and/or PhD applications accepted) |
|
|
Pre-mRNA RNA splicing, adaptation, genetic diseases, cancer, Rett syndrome, (PhD applications accepted) |
Advisors accepting postdoctoral fellows
| Faculty advisor | Research area |
|---|---|
| Jeremy Chopek | Spinal cord physiology; movement; neural circuit formation |
| Sanjiv Dhingra | Cardiac stem cell therapy and tissue engineering, Immunomodulatory biomaterials synthesis, characterization and application, allograft vasculopathy, iPSCs and disease modeling, Inborn metabolic disorders. |
| Ian Dixon | Cardiac fibrosis. Myofibroblast inactivation by endogenous repressor of Smads. Generation of living valve leaflets using matricryptic signals. |
|
Studies to examine the influence of gene/environment interactions on pancreatic islet function and metabolic health: Implications for childhood-onset type 2 diabetes in Indigenous youth |
|
| Soheila Karimi | Translational neuroscience; neural regeneration; neural stem cell biology and therapeutics; multiple sclerosis and spinal cord injury; treatment development; neuroinflammation and immune modulation; myelin repair; organoid; transgenic and genetic models (previous background in neuroscience demonstrated in publication is required) |
|
Interested clinically in minimal invasive pediatric general surgery. Research is focused on congenital anomalies including congenital diaphragmatic hernia (CDH) and abnormal lung development, including delineating the role of microRNAs and circular RNAs |
|
|
Mitochondrial dynamics; cell death; autophagy; myocardial infarction |
|
|
Cancer; genomic instability; nuclear architecture; imaging |
|
| Suresh Mishra | Interplay between adipose and immune functions in normal physiology, aging, and diseases, with a current emphasis on exploring the role of sex hormones, genes, and micro-RNAs in defining sex differences in these functions to improve understanding of sex-biased metabolic and immune diseases. |
| Inna Rabinovich-Nikitin | Circadian factors and pathways by which circadian misalignment leads to cardiometabolic dysfunction; how controlling feeding/fasting cycles affect cardiac function in cardiometabolic syndrome and contribute or prevent the development of cardiometabolic heart failure. |
| Peter Thompson | Type 1 Diabetes; autoimmunity; pancreatic islet biology; cellular senescence; DNA damage response; ER stress. |
|
To study non-alcoholic fatty liver disease (NAFLD) in an obese rodent model. In addition to examining the physiological processes leading to disease onset, a major objective is to identify key cellular mechanisms that may serve as targets for interventions capable of slowing or reversing disease progression. |
News and stories
View more news and stories-

Rady researchers receive early career transition awards
Rady Faculty of Health Sciences
-
Winnipeg Free Press: Satirical musical tackles health-care woes in bite-sized chunks
Rady Faculty of Health Sciences, UM Today
-

The Globe and Mail: 2024 Gairdner Award recipients paved way for revolutions in cancer treatment, genomics, and global health
Rady Faculty of Health Sciences, UM Today
Events
View more eventsYou may also be looking for...
Contact us
Physiology and Pathophysiology
432 Basic Medical Sciences Building
745 Bannatyne Avenue
University of Manitoba
Winnipeg, MB R3E 0J9 Canada